as.data.frame() returns x@data (as data.frame).
as.matrix() returns the spectra matrix x@data$spc as matrix.
as.wide.df() converts the spectra matrix to a data.frame. The extra
data together with this data is returned. The column names of the spectra
matrix are retained (if they are numbers, without preceding letters).
as.long.df() returns a long-format data.frame.
The data.frame returned by as.long.df() is guaranteed to have columns
spc and .wavelength. If nwl(x) == 0 these columns will be NA.
as.t.df() produces a 'transposed' data.frame with columns containing
the spectra.
# S3 method for hyperSpec
as.data.frame(x, row.names = TRUE, optional = NULL, ...)
# S3 method for hyperSpec
as.matrix(x, ...)
as.wide.df(x, wl.prefix = "")
as.long.df(x, rownames = FALSE, wl.factor = FALSE, na.rm = TRUE)
as.t.df(x)Arguments
- x
A
hyperSpecobject.- row.names
If
TRUE, a column.rowis created containing row names or row indices if no rownames are set.If character vector, the rownames are set accordingly.
- optional
(Ignored)
- ...
(Ignored)
- wl.prefix
Prefix to prepend wavelength column names.
- rownames
Should the rownames be in column
.rownamesof the long-format data.frame?- wl.factor
Should the wavelengths be returned as a factor (instead of numeric)?
- na.rm
If
TRUE, rows where spc is notNAare deleted.
Value
as.data.frame()returnsx@dataasdata.frame;as.matrix()returnsx@data$spc(==x$spc==x[[]]) asmatrix;
as.wide.df()returns adata.framethat consists of the extra data and the spectra matrix converted to a data.frame. The spectra matrix is expanded in place.
as.long.df()returns the stacked or molten version ofx@data. The wavelengths are in column.wavelength.
as.t.df()returns a data.frame similar toas.long.df, but each spectrum in its own column. This is useful for exporting summary spectra, see the example.
See also
[[]]for a shortcut toas.matrix()
utils::stack()andreshape::melt()orreshape2::melt()for other functions producing long-format data frames.
Examples
as.data.frame(faux_cell[1:3, , 600 ~ 620])
#> x y region spc.602 spc.606 spc.610 spc.614 spc.618 .row
#> 1 -11.55 -4.77 matrix 15 22 15 13 15 1
#> 2 -10.55 -4.77 matrix 168 166 159 175 160 2
#> 3 -9.55 -4.77 matrix 112 113 99 115 119 3
as.matrix(faux_cell[1:3, , 600 ~ 620])
#> 602 606 610 614 618
#> [1,] 15 22 15 13 15
#> [2,] 168 166 159 175 160
#> [3,] 112 113 99 115 119
lm(c ~ spc, data = flu[, , 450])
#>
#> Call:
#> lm(formula = c ~ spc, data = flu[, , 450])
#>
#> Coefficients:
#> (Intercept) spc
#> 0.0038493 0.0004407
#>
as.wide.df(faux_cell[1:5, , 600 ~ 610])
#> x y region 602 606 610
#> 1 -11.55 -4.77 matrix 15 22 15
#> 2 -10.55 -4.77 matrix 168 166 159
#> 3 -9.55 -4.77 matrix 112 113 99
#> 4 -8.55 -4.77 matrix 23 33 39
#> 5 -7.55 -4.77 matrix 3 0 1
summary(as.wide.df(faux_cell[1:5, , 600 ~ 610]))
#> x y region 602 606
#> Min. :-11.55 Min. :-4.77 cell :0 Min. : 3.0 Min. : 0.0
#> 1st Qu.:-10.55 1st Qu.:-4.77 matrix :5 1st Qu.: 15.0 1st Qu.: 22.0
#> Median : -9.55 Median :-4.77 nucleus:0 Median : 23.0 Median : 33.0
#> Mean : -9.55 Mean :-4.77 Mean : 64.2 Mean : 66.8
#> 3rd Qu.: -8.55 3rd Qu.:-4.77 3rd Qu.:112.0 3rd Qu.:113.0
#> Max. : -7.55 Max. :-4.77 Max. :168.0 Max. :166.0
#> 610
#> Min. : 1.0
#> 1st Qu.: 15.0
#> Median : 39.0
#> Mean : 62.6
#> 3rd Qu.: 99.0
#> Max. :159.0
as.long.df(flu[, , 405 ~ 410])
#> .wavelength spc filename c
#> 1 405.0 27.15000 rawdata/flu1.txt 0.05
#> 2 405.0 66.80133 rawdata/flu2.txt 0.10
#> 3 405.0 93.14433 rawdata/flu3.txt 0.15
#> 4 405.0 130.66367 rawdata/flu4.txt 0.20
#> 5 405.0 167.26667 rawdata/flu5.txt 0.25
#> 6 405.0 198.43033 rawdata/flu6.txt 0.30
#> 1.1 405.5 32.34467 rawdata/flu1.txt 0.05
#> 2.1 405.5 63.71533 rawdata/flu2.txt 0.10
#> 3.1 405.5 103.06767 rawdata/flu3.txt 0.15
#> 4.1 405.5 139.99833 rawdata/flu4.txt 0.20
#> 5.1 405.5 171.89833 rawdata/flu5.txt 0.25
#> 6.1 405.5 209.45800 rawdata/flu6.txt 0.30
#> 1.2 406.0 33.37867 rawdata/flu1.txt 0.05
#> 2.2 406.0 66.71200 rawdata/flu2.txt 0.10
#> 3.2 406.0 106.19367 rawdata/flu3.txt 0.15
#> 4.2 406.0 143.79767 rawdata/flu4.txt 0.20
#> 5.2 406.0 177.47067 rawdata/flu5.txt 0.25
#> 6.2 406.0 215.78500 rawdata/flu6.txt 0.30
#> 1.3 406.5 34.41933 rawdata/flu1.txt 0.05
#> 2.3 406.5 69.58233 rawdata/flu2.txt 0.10
#> 3.3 406.5 110.18633 rawdata/flu3.txt 0.15
#> 4.3 406.5 148.42000 rawdata/flu4.txt 0.20
#> 5.3 406.5 184.62467 rawdata/flu5.txt 0.25
#> 6.3 406.5 224.58700 rawdata/flu6.txt 0.30
#> 1.4 407.0 36.53133 rawdata/flu1.txt 0.05
#> 2.4 407.0 72.52967 rawdata/flu2.txt 0.10
#> 3.4 407.0 113.24867 rawdata/flu3.txt 0.15
#> 4.4 407.0 152.13267 rawdata/flu4.txt 0.20
#> 5.4 407.0 189.75233 rawdata/flu5.txt 0.25
#> 6.4 407.0 232.52800 rawdata/flu6.txt 0.30
#> 1.5 407.5 37.64767 rawdata/flu1.txt 0.05
#> 2.5 407.5 74.55833 rawdata/flu2.txt 0.10
#> 3.5 407.5 119.17300 rawdata/flu3.txt 0.15
#> 4.5 407.5 159.31033 rawdata/flu4.txt 0.20
#> 5.5 407.5 198.11533 rawdata/flu5.txt 0.25
#> 6.5 407.5 240.77133 rawdata/flu6.txt 0.30
#> 1.6 408.0 38.13700 rawdata/flu1.txt 0.05
#> 2.6 408.0 77.04800 rawdata/flu2.txt 0.10
#> 3.6 408.0 121.31333 rawdata/flu3.txt 0.15
#> 4.6 408.0 165.05233 rawdata/flu4.txt 0.20
#> 5.6 408.0 205.56267 rawdata/flu5.txt 0.25
#> 6.6 408.0 248.04667 rawdata/flu6.txt 0.30
#> 1.7 408.5 39.17700 rawdata/flu1.txt 0.05
#> 2.7 408.5 80.25967 rawdata/flu2.txt 0.10
#> 3.7 408.5 124.67533 rawdata/flu3.txt 0.15
#> 4.7 408.5 168.68967 rawdata/flu4.txt 0.20
#> 5.7 408.5 208.41933 rawdata/flu5.txt 0.25
#> 6.7 408.5 256.89133 rawdata/flu6.txt 0.30
#> 1.8 409.0 40.73567 rawdata/flu1.txt 0.05
#> 2.8 409.0 82.53867 rawdata/flu2.txt 0.10
#> 3.8 409.0 129.56867 rawdata/flu3.txt 0.15
#> 4.8 409.0 175.45900 rawdata/flu4.txt 0.20
#> 5.8 409.0 217.55267 rawdata/flu5.txt 0.25
#> 6.8 409.0 262.73900 rawdata/flu6.txt 0.30
#> 1.9 409.5 41.38133 rawdata/flu1.txt 0.05
#> 2.9 409.5 84.49167 rawdata/flu2.txt 0.10
#> 3.9 409.5 134.11733 rawdata/flu3.txt 0.15
#> 4.9 409.5 181.58100 rawdata/flu4.txt 0.20
#> 5.9 409.5 224.74633 rawdata/flu5.txt 0.25
#> 6.9 409.5 270.27133 rawdata/flu6.txt 0.30
#> 1.10 410.0 44.25133 rawdata/flu1.txt 0.05
#> 2.10 410.0 88.15167 rawdata/flu2.txt 0.10
#> 3.10 410.0 139.98667 rawdata/flu3.txt 0.15
#> 4.10 410.0 185.69233 rawdata/flu4.txt 0.20
#> 5.10 410.0 231.03567 rawdata/flu5.txt 0.25
#> 6.10 410.0 281.82867 rawdata/flu6.txt 0.30
summary(as.long.df(flu[, , 405 ~ 410]))
#> .wavelength spc filename c
#> Min. :405.0 Min. : 27.15 Length:66 Min. :0.050
#> 1st Qu.:406.0 1st Qu.: 75.18 Class :character 1st Qu.:0.100
#> Median :407.5 Median :137.05 Mode :character Median :0.175
#> Mean :407.5 Mean :137.80 Mean :0.175
#> 3rd Qu.:409.0 3rd Qu.:196.02 3rd Qu.:0.250
#> Max. :410.0 Max. :281.83 Max. :0.300
summary(as.long.df(flu[, , 405 ~ 410], rownames = TRUE))
#> .rownames .wavelength spc filename c
#> 1:11 Min. :405.0 Min. : 27.15 Length:66 Min. :0.050
#> 2:11 1st Qu.:406.0 1st Qu.: 75.18 Class :character 1st Qu.:0.100
#> 3:11 Median :407.5 Median :137.05 Mode :character Median :0.175
#> 4:11 Mean :407.5 Mean :137.80 Mean :0.175
#> 5:11 3rd Qu.:409.0 3rd Qu.:196.02 3rd Qu.:0.250
#> 6:11 Max. :410.0 Max. :281.83 Max. :0.300
summary(as.long.df(flu[, , 405 ~ 410], wl.factor = TRUE))
#> .wavelength spc filename c
#> 405 : 6 Min. : 27.15 Length:66 Min. :0.050
#> 405.5 : 6 1st Qu.: 75.18 Class :character 1st Qu.:0.100
#> 406 : 6 Median :137.05 Mode :character Median :0.175
#> 406.5 : 6 Mean :137.80 Mean :0.175
#> 407 : 6 3rd Qu.:196.02 3rd Qu.:0.250
#> 407.5 : 6 Max. :281.83 Max. :0.300
#> (Other):30
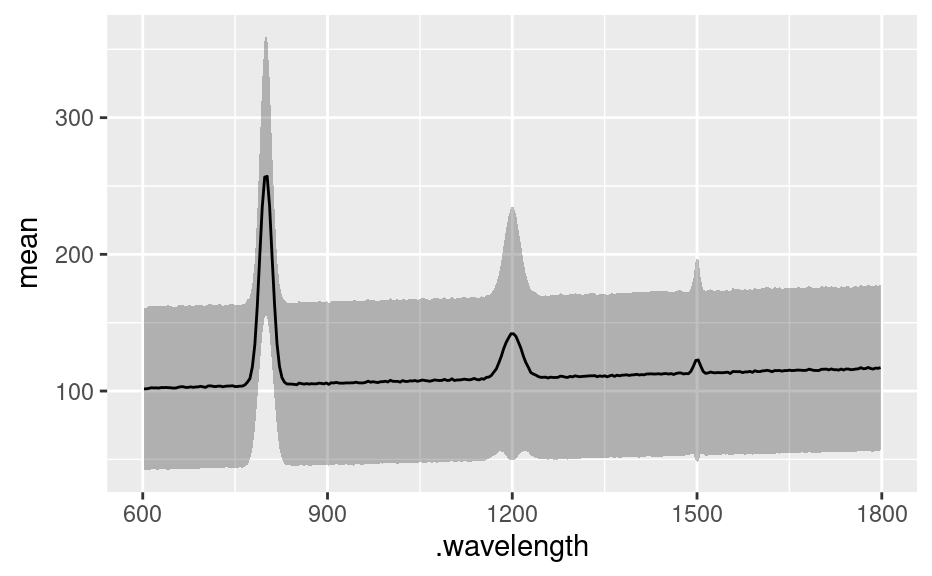
df <- as.t.df(apply(faux_cell, 2, mean_pm_sd))
head(df)
#> .wavelength mean.minus.sd mean mean.plus.sd
#> spc.602 602 42.04492 101.5691 161.0934
#> spc.606 606 42.14042 101.6023 161.0642
#> spc.610 610 41.82034 101.9383 162.0562
#> spc.614 614 42.85471 102.3966 161.9384
#> spc.618 618 42.61228 102.2571 161.9020
#> spc.622 622 42.58845 102.3257 162.0630
if (require(ggplot2)) {
ggplot(df, aes(x = .wavelength)) +
geom_ribbon(aes(ymin = mean.minus.sd, ymax = mean.plus.sd),
fill = "#00000040"
) +
geom_line(aes(y = mean))
}