To create a new hyperSpec object, you can use one of the following functions:
new()(i.e.,new("hyperSpec", ...));hyperSpec().
hyperSpec(
spc = NULL,
data = NULL,
wavelength = NULL,
labels = NULL,
gc = hy_get_option("gc"),
log = "ignored"
)
# S4 method for hyperSpec
initialize(
.Object,
spc = NULL,
data = NULL,
wavelength = NULL,
labels = NULL,
gc = hy_get_option("gc"),
log = "ignored"
)Arguments
- spc
(
matrixor convertible tomatrix)
A spectra matrix with spectra in rows and wavelength intensities in columns.The
spcdoes not need to be an Rmatrix, but it must be an object convertible to a matrix viaI(as.matrix(spc)).- data
(
data.frame)
Adata.framewith extra (non-spectroscopic) data in columns. The data frame may also contain a special columnspcwith amatrixof spectroscopic data. (Such a single column that contains a matrix can be created withdata.frame(spc = I(as.matrix(spc))). However, it is usually more convenient to provide the spectra via thespcargument.)- wavelength
(numeric vector)
The wavelengths corresponding to the columns ofspc.If no wavelengths are given, an appropriate vector is derived from the column names of
data$spc. If this is not possible,1:ncol(data$spc)is used instead.- labels
A named
list:list's element names should contain one or more names of
datacolumns as well as the special name.wavelengthforwavelengths.list's element values should contain the labels for the indicated names, usually in the form of character strings or plotmath expressions. (The labels should be given in a form ready for the text-drawing functions, see
grDevices::plotmath()).
If
labelis not given, a list containingNULLfor each of the columns ofdataandwavelengthis used.- gc
(logical)
Use garbage collection. If the optiongcis set toTRUE, the initialization will have frequent calls tobase::gc(), which can help avoid swapping or running out of memory. The default value ofgccan be set viahy_set_options().- log
This parameter is currently ignored. It is present due to backward compatibility.
- .Object
A new
hyperSpecobject.
Note
A hyperSpec object is an S4 object, so its initialization is performed
by calling new("hyperSpec", ...). The function hyperSpec() is provided
for convenience.
See also
methods::new()for more information on creating and initializing S4 objects.grDevices::plotmath()for expressions used for math annotations as in thelabelslot.hy_set_options()for settinghyperSpecoptions.
Examples
new("hyperSpec")
#> hyperSpec object
#> 0 spectra
#> 1 data columns
#> 0 data points / spectrum
hyperSpec()
#> hyperSpec object
#> 0 spectra
#> 1 data columns
#> 0 data points / spectrum
spc <- matrix(rnorm(12), ncol = 4)
new("hyperSpec", spc = spc)
#> hyperSpec object
#> 3 spectra
#> 1 data columns
#> 4 data points / spectrum
hyperSpec(spc = spc)
#> hyperSpec object
#> 3 spectra
#> 1 data columns
#> 4 data points / spectrum
new("hyperSpec",
data = data.frame(x = letters[1:3]),
spc = spc
)
#> hyperSpec object
#> 3 spectra
#> 2 data columns
#> 4 data points / spectrum
colnames(spc) <- 600:603
new("hyperSpec", spc = spc) # wavelength taken from colnames (spc)
#> hyperSpec object
#> 3 spectra
#> 1 data columns
#> 4 data points / spectrum
# given wavelengths take precedence over colnames of spc
new("hyperSpec", spc = spc, wavelength = 700:703)
#> hyperSpec object
#> 3 spectra
#> 1 data columns
#> 4 data points / spectrum
# specifying labels
h <- new("hyperSpec",
spc = spc, data = data.frame(pos = 1:3),
label = list(
spc = "I / a.u.",
.wavelength = expression(tilde(nu) / cm^-1),
pos = expression("/"(x, mu * m))
)
)
plot(h)
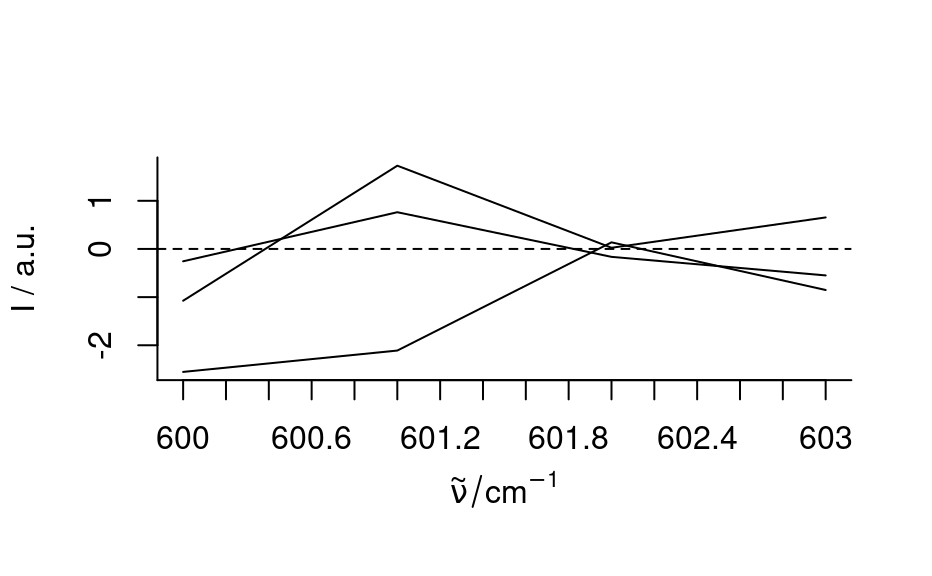 plot_c(h, spc ~ pos)
#> Warning: Intensity at first wavelengh only is used.
plot_c(h, spc ~ pos)
#> Warning: Intensity at first wavelengh only is used.
