The plot method for hyperSpec objects is a switchyard to plot_spc(),
plot_map(), and plot_c(). The function also supplies some convenient
abbreviations for frequently used plots (see 'Details').
# S4 method for hyperSpec,missing
plot(x, y, ...)
# S4 method for hyperSpec,character
plot(x, y, ...)Arguments
- x
hyperSpecobject.- y
String (
"spc","map", etc.) to select what type of plot should be produced. See section 'Details' for available values. Ifyis missing,plot(x)behaves likeplot(x, y = "spc").- ...
Arguments passed to the respective plot function
Details
Supported values for y are:
- "spc" or nothing
calls
plot_spc()to produce a spectra plot.- "spcmeansd"
plots mean spectrum +/- one standard deviation
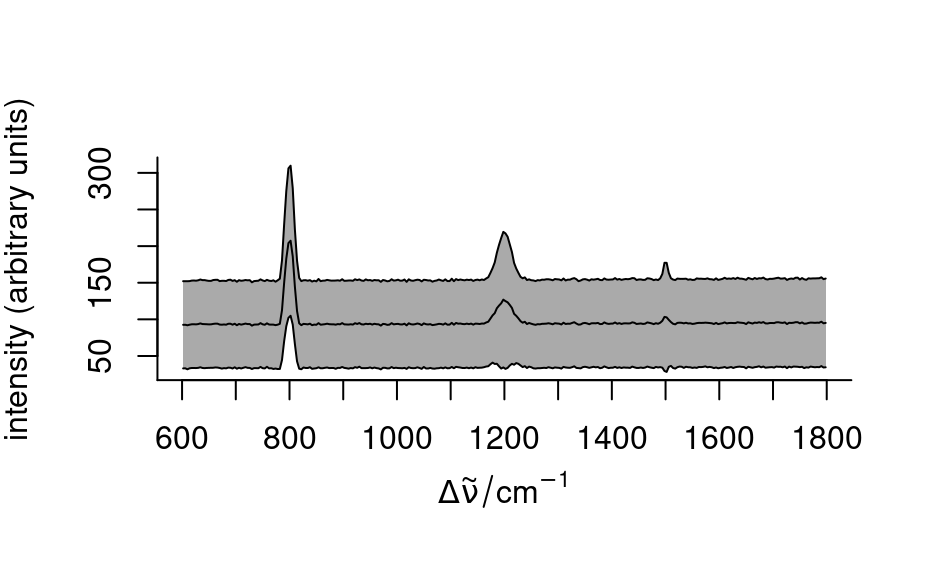
- "spcprctile"
plots 16th, 50th, and 84th percentile spectra. If the distributions of the intensities at all wavelengths were normal, this would correspond to
"spcmeansd". However, this is frequently not the case. Then"spcprctile"gives a better impression of the spectral data set.- "spcprctl5"
like
"spcprctile", but additionally the 5th and 95th percentile spectra are plotted.- "map"
calls
plot_map()to produce a map plot.- "voronoi"
calls
plot_voronoi()to produce a Voronoi plot (tessellated plot, like "map" for hyperSpec objects with uneven/non-rectangular grid).- "mat"
calls
plot_matrix()to produce a plot of the spectra matrix (not to be confused withgraphics::matplot()).- "c"
calls
plot_c()to produce a calibration (or time series, depth-profile, or the like).- "ts"
plots a time series: abbreviation for
plot_c(x, use.c = "t").- "depth"
plots a depth profile: abbreviation for
plot_c(x, use.c = "z").
See also
plot_spc() for spectra plots (intensity over wavelength),
plot_map() for plotting maps, i.e. color coded summary value on two
(usually spatial) dimensions.
Examples
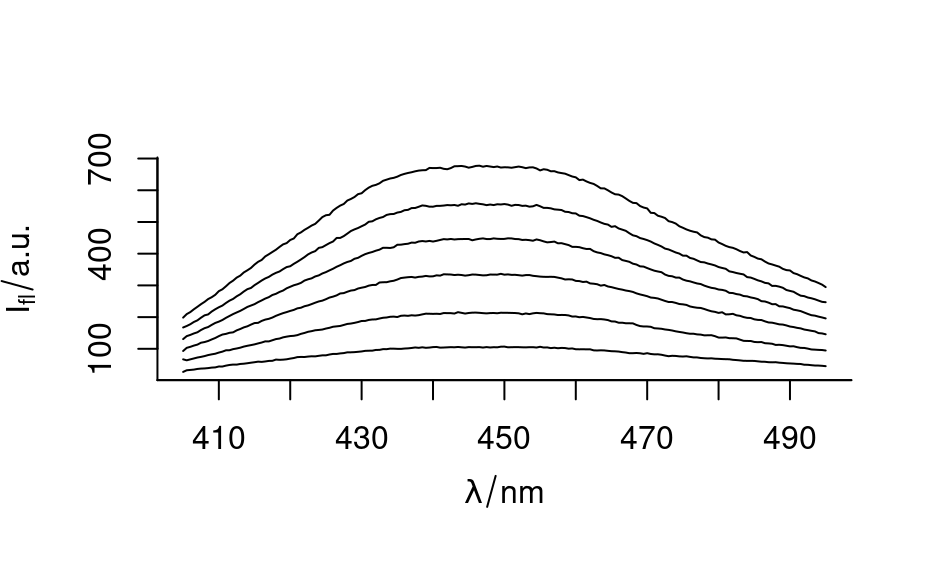
plot(flu)
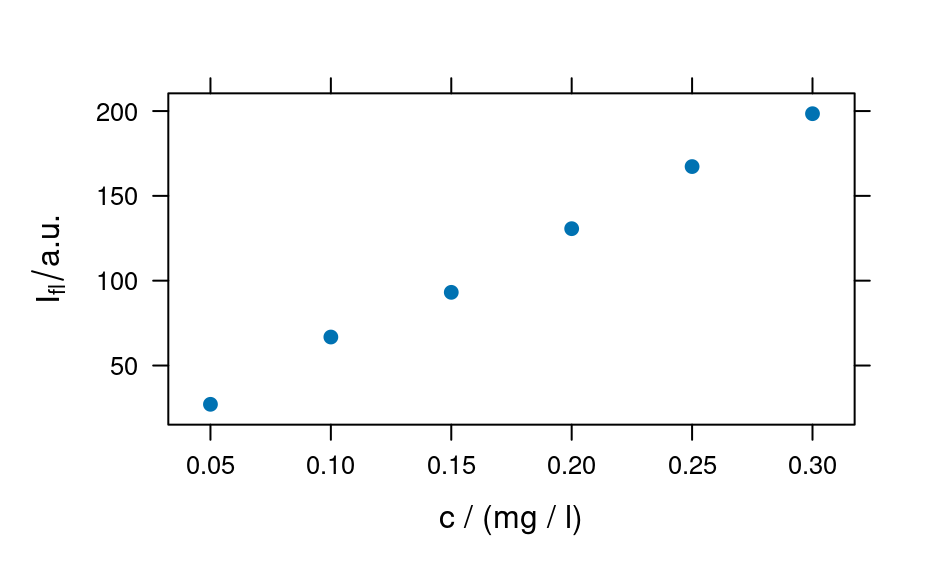
 plot(flu, "c")
#> Warning: Intensity at first wavelengh only is used.
plot(flu, "c")
#> Warning: Intensity at first wavelengh only is used.
 plot(laser, "ts")
#> Warning: Intensity at first wavelengh only is used.
plot(laser, "ts")
#> Warning: Intensity at first wavelengh only is used.
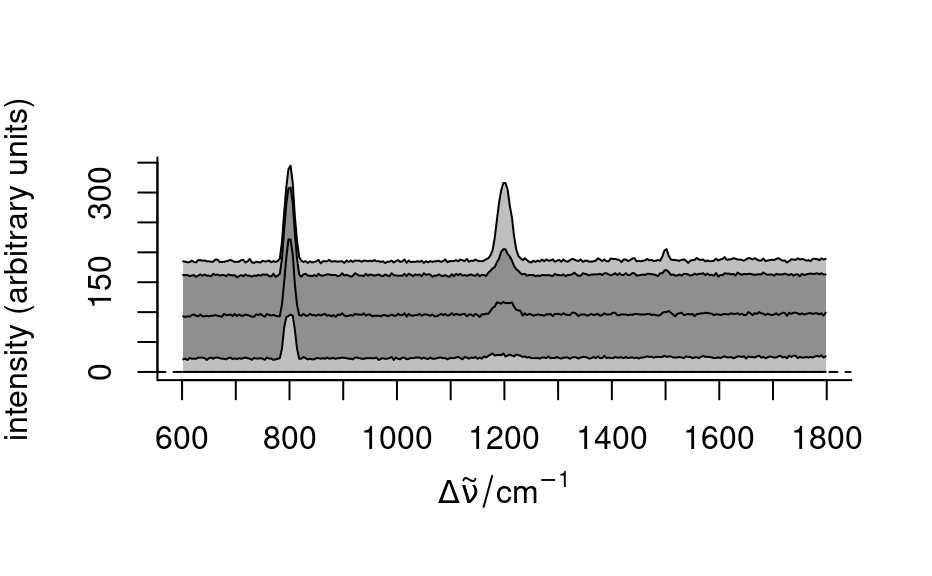
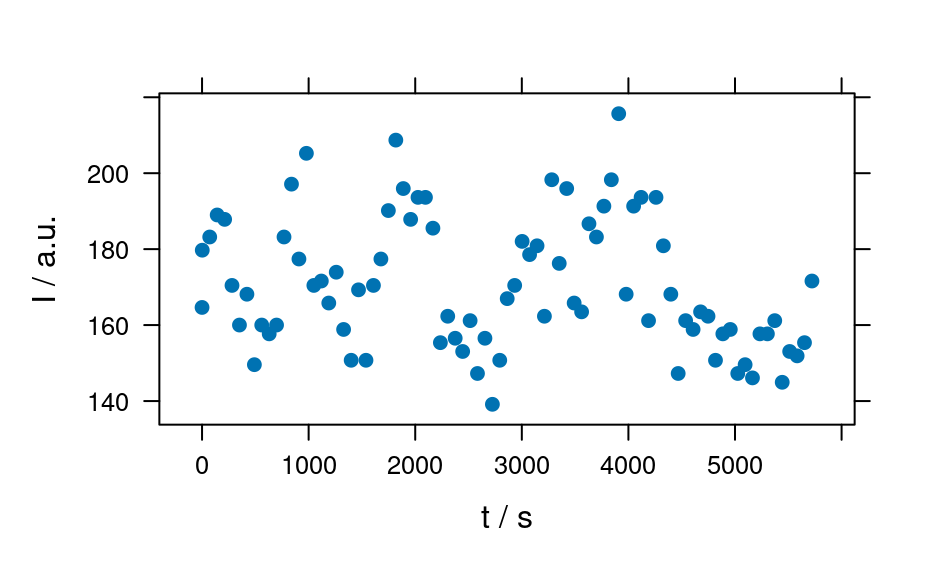 spc <- apply(faux_cell, 2, quantile, probs = 0.05)
spc <- sweep(faux_cell, 2, spc, "-")
plot(spc, "spcprctl5")
spc <- apply(faux_cell, 2, quantile, probs = 0.05)
spc <- sweep(faux_cell, 2, spc, "-")
plot(spc, "spcprctl5")
 plot(spc, "spcprctile")
plot(spc, "spcprctile")
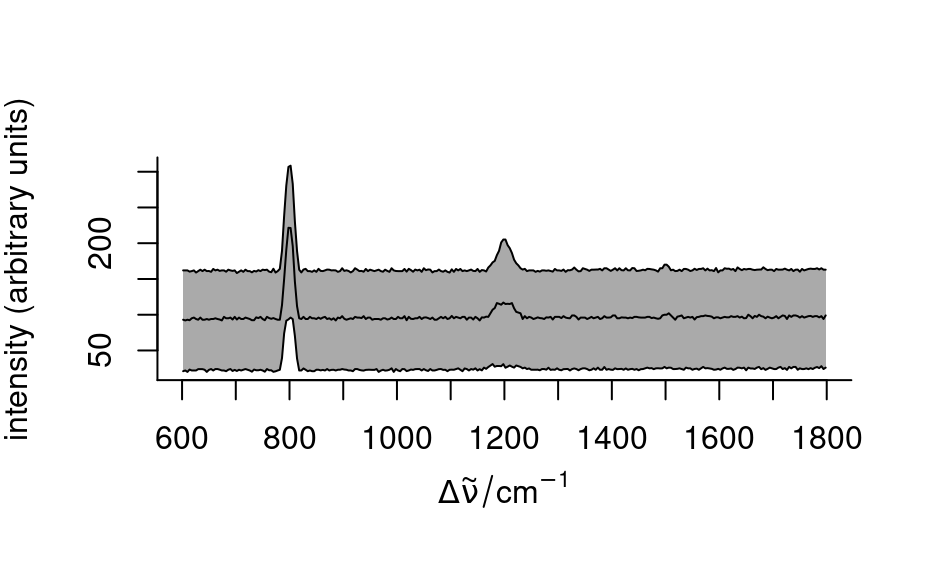 plot(spc, "spcmeansd")
plot(spc, "spcmeansd")
 ### Use plot_spc() as a default plot function.
### Use plot_spc() as a default plot function.